News
 Opal 4.2 Released
Opal 4.2 Released
We are pleased to announce that Opal 4.2 is now available. Opal is OBiBa’s core data management application for biobanks.
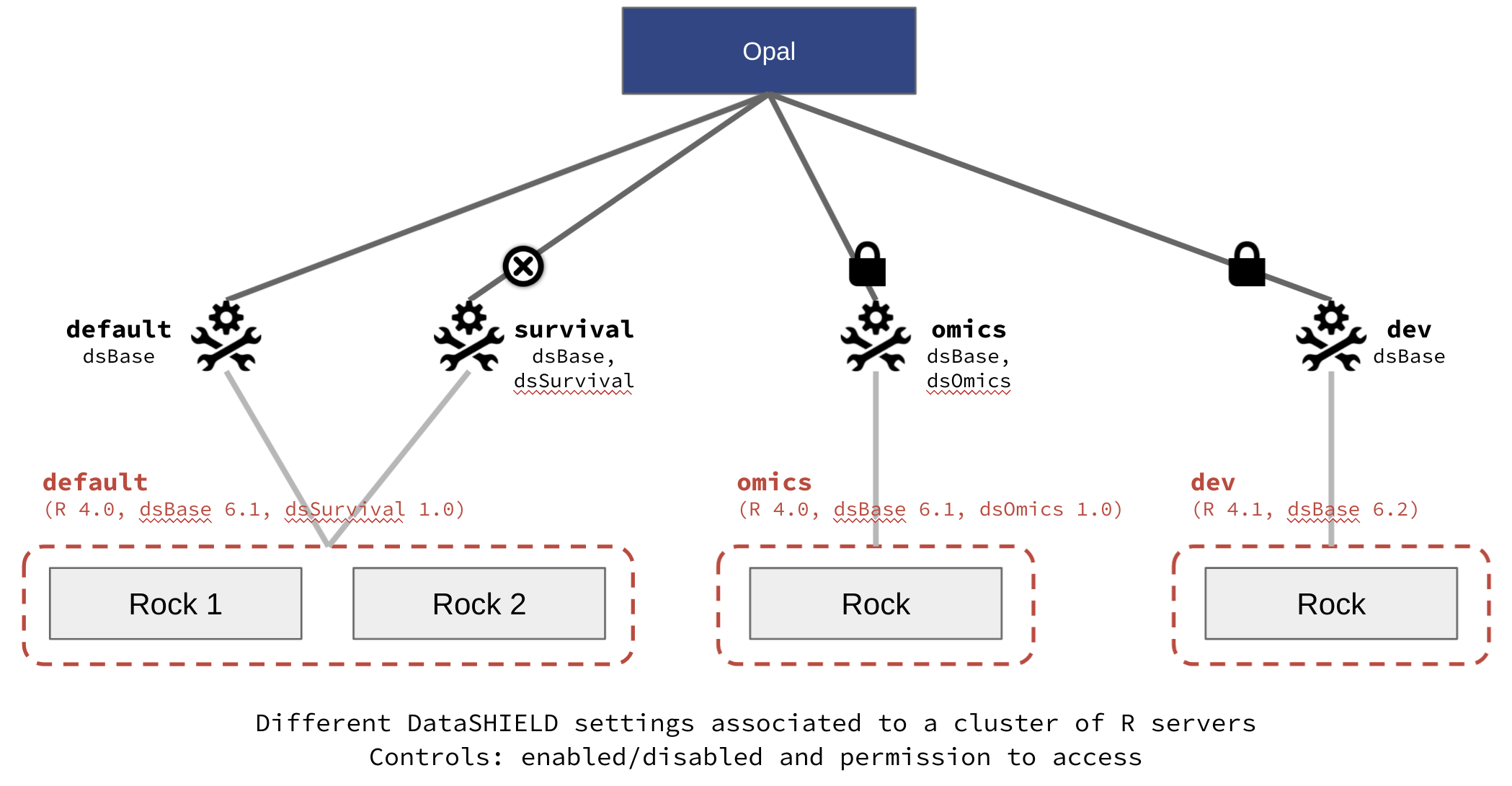
This release of Opal now supports multiple clusters of Rock R servers. This allows to have different R environments: different R versions and different R packages installed.

This fully integrates with the DataSHIELD federated analysis platform by introducing DataSHIELD profiles:
- profile links to a specific R execution environment (R version, R packages installed),
- profile can be enabled/disabled,
- profile usage can be restricted to users with proper permissions,
- profile configuration (allowed aggregation/assignment functions, R options) can be amended,
- profile can be associated to a specific DataSHIELD R parser version.
Additional changes:
- OpenID Connect providers username and groups extraction process can be configured.
- variable annotation user interface can manage taxonomies with thousands of terms.
- user with table/view addition permission can now remove the tables/views created and update data dictionaries in batch.
- operations are faster thanks to an improved permissions cache.
- DataSHIELD R parser updated by DataSHIELD team with new rules: subset syntax (using
[]) is not permitted anymore.
Breaking changes
This Opal release is backward compatible: no major breaking changes are expected.
- DataSHIELD users may encounter some issues because the new DataSHIELD R parser (designed by the DataSHIELD team) is a bit more restrictive. Contact the DataSHIELD team for an alternative to the subset syntax (or apply former DataSHIELD R parser version to the DataSHIELD profile).
R and DataSHIELD packages
These packages have been updated and are backward compatible:
# opal R client, with profiles support
install.packages("opalr")
# DataSHIELD interface, with profiles support
install.packages("DSI")
# DataSHIELD interface Opal implementation
install.packages("DSOpal")
Documentation
See Opal documentation for installation and operation instructions. We recommend to have a look at the section about Docker Image Installation .
See also Opal R package documentation which contains useful articles for managing Opal system and data from R.
Aknowledgements
This release was possible thanks to the funding of the EUCAN-Connect program.